
TNM staging towards a personalized approach in metastatic urothelial carcinoma: what will the future be like?—a narrative review
Introduction
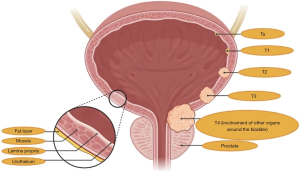
Urothelial carcinoma (UC) is among the most common malignancies worldwide, with around 450,000 new diagnoses each year (1). To date, UC represents the eleventh most frequently diagnosed malignancy in both sexes and the seventh in the male population throughout the world (2). The most important risk factor is tobacco smoking, which is held responsible for the 50% of all UCs, followed by pelvic radiation, occupational exposure to carcinogens and genetic predisposition (3). Overall, about three quarters of UC patients are diagnosed with non-muscle-invasive bladder cancer and are treated with transurethral resection and intravesical instillation of Bacillus of Calmette-Guerin (BCG) or other anticancer agents (4,5). Unfortunately, the remaining 25% of patients presents with muscle-invasive bladder cancer (MIBC)—classically defined by the invasion of the detrusor muscle—or metastatic disease (6). Moreover, approximately the 50% of patients with tumor stages among T2b and T4a develop metastatic disease following radical surgery (Figure 1) (7). Despite platinum-based chemotherapy is the backbone of treatment for metastatic UC, with the combination of cisplatin plus gemcitabine or methotrexate, vinblastine, doxorubicin and cisplatin (M-VAC) (8,9), an important percentage of patients is unfit to receive cisplatin because of comorbidities, old age, peripheral neuropathy and/or poor Eastern Cooperative Oncology Group performance status (ECOG-PS) (10).
The last decade has witnessed notable advances in UC management and previous treatment paradigms of metastatic disease have been modified by immune checkpoint inhibitors (ICIs), which have rapidly emerged as novel therapeutic options (11). In fact, although platinum-based regimens remain the standard first-line treatment for cisplatin-eligible advanced UC patients, therapeutic options are dramatically evolving with the US Food and Drug Administration (FDA) approval of second-line pembrolizumab, nivolumab, avelumab, atezolizumab and durvalumab (12). Moreover, the therapeutic algorithm of metastatic UC is further evolving, with the results of the JAVELIN Bladder 100 trial which have been recently presented at the virtual 2020 ASCO Annual Meeting (13). This phase III trial comparing maintenance avelumab versus best supportive care in UC patients who achieved stable disease, partial response or complete response from first-line platinum-based chemotherapy, has reported a significant improvement in overall survival (OS) in the avelumab arm (21.4 versus 14.3 months; HR, 0.69; 95% CI, 0.56–0.86; P<0.001). Thus, first-line maintenance therapy with the anti-PD-L1 avelumab is destined to become a new standard of care in patients with advanced UC achieving disease control with first-line platinum-based chemotherapy. Nonetheless, several issues remain since the prognosis of patients affected by metastatic disease is still dismal, with a 5-year OS of around 10% (14).
If the American Joint Committee of Cancer (AJCC) tumor-node-metastasis (TNM) classification has among its purposes to properly define cancer staging, the identification of different UC molecular features has allowed to integrate the TNM model with brand-new elements (Table 1) (15,16). In fact, the integration between TNM anatomic-based characteristics, baseline clinical features and the molecular landscape of UC has led to a novel, personalized paradigm in cancer management (17). From a molecular point of view, UC resulted to be a heterogeneous disease, with high mutational rate and genomic instability. In fact, UC is characterized by a marked inter-tumoral and intra-tumoral heterogeneity, which have contributed to the lack of effective targeted treatments in early studies (18). In the last decade, the molecular landscape of UC has begun to emerge, offering the possibility to unveil the basis of UC carcinogenesis and tumor progression (19); moreover, molecular profiling of UCs has become increasingly meaningful due to the identification of potentially targetable molecular alterations, including Fibroblast Growth Factor Receptor (FGFR), Human Epidermal Growth Factor Receptors and DNA damage response (DDR) pathway (20,21). In fact, multiplatform genomic profiling has paved the way towards a new era in UC management, where biomarker-driven clinical trials appear as a mandatory need.

Full table
In the current review, we discuss recent advances regarding the characterization of UC, having the potential to integrate the TNM classification with molecular subtyping in this aggressive disease. A comprehensive literature search on PubMed/Medline, Cochrane library and Scopus has been performed using the keywords “bladder cancer” OR “bladder carcinoma” OR “urothelial carcinoma” OR “muscle-invasive bladder cancer” AND “gene signatures” OR “TCGA” OR “genomic subtypes”. We selected the most relevant and pertinent reports on the basis of the quality of the studies in terms of their applicability, how they were conducted and the number of patients included. Despite molecular profiling studies have better defined the genetic landscape of UC, suggesting the presence of different patterns of mutations, advanced/metastatic UC remains a complex, difficult-to-treat malignancy and more work is warranted in the near future in this direction. We present the following article in accordance with the NARRATIVE REVIEW reporting checklist (available at http://dx.doi.org/10.21037/tau-20-1109).
NGS and baseline characteristics in clinical practice: what we should remember
After the completion of the Human Genome Project in 2003, sequencing of cancer genomes has represented one of the hottest topics in cancer research, with a view to led to a better comprehension of the genetic basis of oncogenesis and tumor progression (22). Concurrently, next-generation sequencing (NGS) technology has incredibly expanded with notable advances in terms of reliability, data interpretation and costs, making NGS feasible in everyday clinical practice (23,24). Before NGS, molecular tumor alterations were identified using single gene assays; conversely, NGS technology has allowed to perform simultaneous analyses of hundreds of genes through targeted sequencing panels (25). In fact, NGS has allowed a faster and simpler sequencing, improving clinicians and researchers understanding of cancer and promoting the birth of a new era, that of precision medicine—and precision oncology (26). One of the aims of precision oncology is to tailor oncological treatments to the single patient’s characteristics, on the basis of a deep characterization, the identification of druggable mutations and the presence of specific biomarkers (27). And importantly, the use of NGS has created the basis for a new horizon, moving from an anatomical and clinical “stratification”—according to clinical, anatomical and pathological features—to a more personalized approach (28).
As we shall see later, UC is a heterogenous, complex disease including several tumor subtypes presenting important and emerging differences, which have been only partially identified. In the efforts towards precision oncology, NGS is a golden tool which is assuming and will assume an increasingly important role; in fact, incorporating genomic information in the diagnostic and staging processes is one of the current and future challenges in UC management, a crucial step in order to improve clinical outcomes in this aggressive disease (29). In this regard, several research groups have harnessed the use of NGS technology to reveal the complex and heterogeneous molecular landscape of UC (30).
Nonetheless, the definition of tumor histology, anatomic-based TNM stage and genomic features is not enough. Traditionally, despite platinum-based chemotherapy with cisplatin plus gemcitabine or M-VAC has represented and still represents the standard first-line treatment in metastatic disease, the use of these regimens is widely limited due to related toxicities and patient comorbidities (31). More specifically, around the 50% of metastatic UC are not eligible for cisplatin-based regimen, with cisplatin ineligibility usually defined as follows: ECOG-PS ≥2 and/or creatinine clearance <60 mL/min and/or hearing loss of 25 dB at 2 contiguous frequencies and/or peripheral neuropathy grade ≥2 and/or New York Heart Association class ≥3 heart failure (32). Moreover, age is another important element limiting the use of cisplatin in UC patients (32).
The recent advent of ICIs has changed the front-line setting of cisplatin-ineligible patients, with two trials showing that atezolizumab and pembrolizumab have been suggested to be feasible and effective strategies (33)—we will not discuss the details regarding the trials assessing these and other PD-1 and PD-L1 inhibitors in this setting, that are beyond the scope of this paper. Nonetheless, immunotherapy has its caveats. T-cell activation caused by ICIs can be responsible of immune-related adverse events (irAEs), including skin reactions, thyroid dysfunction, pneumonitis, hepatitis and other toxicities that usually do not occur with conventional cytotoxic chemotherapy (34). More specifically, the incidence of any grade irAEs has been reported to range from 40% to 60% in patients receiving anti-PD-1 and anti-PD-L1 agents, and since irAEs are different from adverse events of systemic chemotherapy, these are frequently underestimated and even not detected (35,36). Lastly, irAEs may led to the necessity of frequent monitoring, to the use of immunosuppressive therapies, and sometimes, to hospitalization and death (37).
Modern oncology has seen a passage from an organ-centric, anatomic-based vision to a deep molecular analysis, moving towards a personalized approach. All things considered, although genomic studies have opened the door of new world, personalized oncology cannot overlook clinical features and underlying comorbidities, two elements which are and remain the mainstay of treatment choices in UC—today as yesterday.
UC: the molecular landscape
The genomic characterization of UC has suggested the presence of different biological subtypes of disease, with a diverse mutational landscape (38). Early reports evidenced the presence of two major groups mimicking the breast cancer classification—luminal and basal, which corresponded to different stages in urothelial differentiation (Figure 2) (39). Luminal subgroup was reported to express high levels of low molecular weight keratin 20, PPARG, FGFR3 and uroplakins while the basal subgroup was associated with high levels of EGFR, CD44 and high molecular weight keratins, including keratin 14 and 15. Interestingly, it has been hypothesized that basal cells could present important analogies with triple-negative breast cancer cells, which similarly express high molecular weight keratins and stem cell markers such as CD44 (40). Further studies have subsequently identified more molecular subtypes of UC, according to the expression of examined genes (Figure 2).
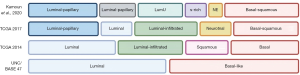
The Cancer Genome Atlas (TCGA) project for bladder cancer had the merit to shed light on this complex and underground landscape, with two major studies focusing on DNA, RNA and protein analyses (41,42). The first TCGA study included 131 bladder cancer patients, where the integrated genomic analysis showed high somatic mutation rate (median 5.5/Megabase) and 32 significant gene mutations (41). Moreover, the 69% of bladder cancers presented genomic alterations which in the 44% of cases concerned the receptor tyrosine kinase/MAPK pathway and in the 42% the PI3K/AKT/mTOR, according to the results of this analysis. Interestingly, several alterations in the receptor tyrosine kinase/RAS were identified such as FGFR3 activations, EGFR amplifications, ERBB2 and ERBB3 mutations. This first TCGA report described four cancer subtypes: luminal, luminal infiltrated, basal and squamous (Figure 2).
In 2017, the results of the TCGA expanded cohort analysis on 412 chemotherapy-naïve samples of MIBC cases have been published, confirming the high mutation rate which characterizes this malignancy (42); moreover, this report detected 64 significant mutated genes, a higher number compared with the 32 mutations of the 2014 analysis. Interestingly, the 412 MIBCs were split in 5 different expression subtypes according to RNA expression analysis: luminal-papillary (35%), luminal (6%), basal-squamous (35%), luminal-infiltrated (19%) and neuronal (5%) (42), a classification which was also proposed in a view to stratify patients for specific therapeutic options (Figure 2). Lastly, the authors defined 4 major groups on the basis of distinct mutational signatures (42).
More recently, an international consensus proposed a MIBC classification on the basis of 1,750 transcriptomic profiles from 18 databases (43). According to this classification, a consensus set of six molecular classes has been defined: luminal papillary (24%), luminal nonspecified (8%), luminal unstable (15%), stroma-rich (15%), basal-squamous (35%) and neuroendocrine-like (3%) (Figure 2). Interestingly, these classes differ according to infiltration by immune and stromal cells, oncogenic mechanisms, histological and clinical features, suggesting possible therapeutic implications. For instance, the luminal papillary class showed high rate of FGFR3 mutations and translocations, suggesting that FGFR inhibitors could represent effective treatments in these patients. Conversely, the luminal unstable class presented high rate (76%) of TP53 mutations, the basal-squamous high expression of basal differentiation markers and the neuroendocrine-like subgroup inactivation of TP53 and RB1. According to this report by Kamoun and colleagues, stroma-rich tumors and luminal papillary malignancies had the best prognosis while neuroendocrine-like and basal-squamous tumors presented worse prognosis (43).
Molecular testing and treatment choices
Although the identification of molecular subtypes has the potential to guide disease management and therapeutic choices, prognostic and clinical implications of UC subtypes remain largely unclear (44). Importantly, targeting UCs on the basis of molecular subtypes is a very challenging option, given the not negligible heterogeneities and methodological issues (19). At the same time, an increasing emphasis has been recently placed on potential predictive biomarkers, including FGFR alterations, PD-L1 expression, DDR genes, and their relationship with molecular subtypes (45).
The FGFR pathway has been involved in the modulation of several biological processes such as cell survival, proliferation, differentiation and angiogenesis (46). As in the case of other malignancies, the four transmembrane receptor tyrosine kinases FGFR1, FGFR2, FGFR3 and FGFR4 can present molecular alterations in UC and, on the basis of the physiological activity of FGFR, aberrations of this signaling may play an important role as pro-oncogenic drivers (47). The frequency of FGFR3 mutations in MIBC is reported to be less than 25% while activating point mutations are more common in early-stage disease (48). In terms of molecular classes, recent reports have highlighted that the luminal subgroup is associated with lower CD8+ genes, higher FGFR3 expression and resistance to ICIs (49). Interestingly, these data suggest that patients with FGFR3 aberrations might not benefit from immunotherapy, thus guiding therapeutic choice towards FGFR targeted therapies. An exploratory analysis of IMvigor210 trial reported that luminal I subtype patients had lower PD-L1 immune cell expression and CD8+ genes expression, thus achieving lower response rates to the anti-PD-L1 atezolizumab (50). Conversely, PD-L1 expression on immunohistochemistry resulted high in the basal subtype, where enriched PD-L1 expression was not related with ORR to the PD-L1 inhibitor, which was in turn significantly higher in luminal cluster II (ORR 34%). Overall, these findings contrast with a similar analysis of the CheckMate275 trial, where patients belonging to the TCGA basal subtype had highest response rate to nivolumab (51). It is worth noting that both studies analyzed biopsies from different specimens for the TCGA analysis, including metastatic lesions, nodal sites of disease and primary tumors. Moreover, the lack of standardization of the TCGA classification in stratifying patients according to molecular subtypes could have represented an important source of bias. Consequently, strong evidence-based conclusions regarding the real impact of TCGA subtyping as a predictive biomarker for ICIs response cannot be drawn so far.
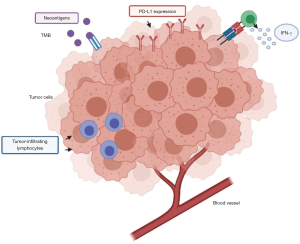
As stated above, although ICIs have shown clinical activity in advanced UC, modifying the therapeutic scenario in this setting, an important percentage of patients does not receive any benefit from immunotherapy due to fast disease progression and lack of response (52). Therefore, biomarkers able to predict response to ICIs would be needed (53). Nonetheless, although some potential associations between biomarkers and responses to immunotherapy have been suggested, these biomarkers have not yet been validated. In terms of predictors of response to ICIs, PD-L1 expression, tumor infiltrating CD8+ lymphocytes and tumor mutational burden (TMB) have been widely studied in several malignancies, including UC (Figure 3) (54). With regard to PD-L1 expression, it is worth noting that there is no standardized format to assess PD-L1 with immunohistochemistry (55). Moreover, thresholds to define PD-L1 positivity vary in different trials and methods themselves of evaluation of PD-L1 may be based on immunohistochemistry or tumor cells (56). Overall, the presence of different assays and scoring systems to define the cut-off positivity for PD-L1 expression is source of confusion, with different trials reporting conflicting results. Probably, the use of a single, standardized method to assess PD-L1 positivity would be the first step to follow, before trying to establish its predictive value in UC patients receiving ICIs.
Another potential predictive biomarker is TMB—commonly defined as the overall number of mutations detected in cancer cells (57). Recent reports have associated increased TMB with more favorable responses to ICIs and, according to results described from the TCGA project, UC presents the third highest mutation rate after melanoma and lung cancer (41). Data from an IMvigor210 subgroup analysis suggested that higher TMB could correlate with clinical benefit, with higher ORR and longer OS in patients receiving atezolizumab (50). Interestingly, median mutation load was 6.4 mut/Megabase in subjects that non responded to atezolizumab compared with 12.4 median mut/Megabase in atezolizumab-responders. Nonetheless, a subsequent reanalysis by whole-exome sequencing did not confirm this association between TMB and response to atezolizumab (58). Lastly, another report from the CheckMate275 trial suggested a correlation between high TMB and better PFS (3.02 versus 1.87 months) and ORR (31.9% versus 17.4%) (51). Prospective data on larger cohorts of patients are still necessary to clarify the role of TMB in UC, a biomarker which undoubtedly needs further validation.
High levels of CD8+ T cells characterize the T cell tumoral inflammation, together with an increased interferon (IFN) and TH1-like chemokine expression (59). When T cells are activated, these are able to proliferate and to differentiate, with subsequent release of pro-inflammatory cytokines (Figure 3) (60). These cytokines include IFN-γ, leading to an upregulation of PD-L1 and PD-L2 (61); on the basis of PD-L1 protein expression and the number of tumor-infiltrating lymphocytes (TILs), the tumor microenvironment (TME) is usually defined as non-immunogenic (“cold”) or immunogenic (“hot”), and the assessment of TME immunogenicity has been suggested as a useful guide for treatment decision (62). For instance, high levels of IFN-γ and higher density of TILs were associated with increased ORR to atezolizumab in the IMvigor210 trial; similarly, 177 tumor samples correlated to responses to the anti-PD-1 agent nivolumab showed a higher IFN-γ signature in CheckMate275 (50,51). However, these data are still preliminary and further studies are needed since ICIs-responders in these two trials were not only patients with inflamed cytokine signatures.
Other interesting biomarkers are DDR genes alterations, which have been associated with a reduced ability to repair DNA damage (63). In fact, the role of DDR genes consists in maintaining genomic stability, repairing DNA damages (64), and in physiological conditions, the mechanism repairs damaged nucleotides without harmful effects; conversely, in presence of DDR genes alterations, repair mechanisms cannot work, resulting in genomic instability (65). More specifically, the DDR pathways are able to recognize DNA damage, to stop cell cycle and to play a fundamental role in DNA repair (66)—where a key element is represented by the Poly (ADP-ribose) Polymerase 1 and 2 (PARP1 and PARP2) genes and whose inhibition surely represents one of the hottest topics in current cancer research (67). Around the 38% of MIBC patients has been reported to present mutations in genes involved in the DDR pathway and previous reports have suggested that DDR gene alterations could play a prognostic role in metastatic UC (68); nonetheless, this prognostic role is still to be clarified. In fact, while some reports suggested that tumors with low excision repair cross complementing 1 (ERCC1) mRNA expression could be associated with longer survival (69), other studies have highlighted worse survival in patients with high expression of ERCC1, RAD51 and PAR at immunohistochemistry (70). The prolonged survival of patients harboring DDR genes mutations appears intimately linked to the sensitivity of platinum-based chemotherapy, as previously found in other malignancies (e.g., ovarian cancer, breast cancer and pancreatic adenocarcinoma) (71,72). Lastly, higher mutational load and TILs have been identified in patients with DDR gene alterations, thus providing the rationale for the testing of ICIs in this setting (73). Interestingly, a recent retrospective study has detected a statistically significant association between DDR alterations and response to PD-1 and PD-L1 inhibitors in metastatic UC patients receiving nivolumab or atezolizumab (74). Further efforts are warranted in this direction in order to better define if alterations in DDR genes could represent a potential marker of clinical benefit in patients treated with modern immunotherapy.
Conclusions
Despite notable advances in the understanding of molecular features characterizing this disease, metastatic UC remains a difficult to treat malignancy, and therapy is still palliative. A broad range of recent studies have suggested the presence of UC molecular subtypes, the characteristics and therapeutic implications of whom still need to be clarified. In this changing landscape, more efforts are needed to identify UC patients who are most likely to benefit from medical treatment, whether it is ICIs, targeted therapies or other novel emerging drugs, through a 360-degree evaluation including clinical, anatomic and molecular features.
Acknowledgments
Funding: None.
Footnote
Provenance and Peer Review: This article was commissioned by the Guest Editors (Rodolfo Montironi, Alessia Cimadamore, Antonio Lopez-Beltran, Marina Scarpelli and Liang Cheng) for the series “Update on Molecular Classification and Individualized Treatments of Genitourinary Tumors” published in Translational Andrology and Urology. The article was sent for external peer review organized by the Guest Editor and the editorial office.
Reporting Checklist: The authors have completed the NARRATIVE REVIEW reporting checklist. Available at http://dx.doi.org/10.21037/tau-20-1109
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tau-20-1109). The series “Update on Molecular Classification and Individualized Treatments of Genitourinary Tumors” was commissioned by the editorial office without any funding or sponsorship. AC, MS, LC, ALB and RM served as the unpaid Guest Editors of the series. LC serves as an unpaid editorial board member of Translational Andrology and Urology from Dec 2018 to Nov 2022. The authors have no other conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Antoni S, Ferlay J, Soerjomataram I, et al. Bladder cancer incidence and mortality: a global overview and recent trends. Eur Urol 2017;71:96-108. [Crossref] [PubMed]
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin 2019;69:7-34. [Crossref] [PubMed]
- Burger M, Catto JW, Dalbagni G, et al. Epidemiology and risk factors of urothelial bladder cancer. Eur Urol 2013;63:234-41. [Crossref] [PubMed]
- El-Achkar A, Souhami L, Kassouf W. Bladder preservation therapy: review of literature and future directions of trimodal therapy. Curr Urol Rep 2018;19:108. [Crossref] [PubMed]
- Smith AB, Deal AM, Woods ME, et al. Muscle-invasive bladder cancer: evaluating treatment and survival in the National Cancer Data Base. BJU Int 2014;114:719-26. [Crossref] [PubMed]
- Donat SM, Shabsigh A, Savage C, et al. Potential impact of postoperative early complications on the timing of adjuvant chemotherapy in patients undergoing radical cystectomy: a high-volume tertiary cancer center experience. Eur Urol 2009;55:177-85. [Crossref] [PubMed]
- Ploussard G, Shariat SF, Dragomir A, et al. Conditional survival after radical cystectomy for bladder cancer: evidence for a patient changing risk profile over time. Eur Urol 2014;66:361-70. [Crossref] [PubMed]
- Sternberg CN, de Mulder P, Schornagel JH, et al. Seven year update of an EORTC phase III trial of high-dose intensity M-VAC chemotherapy and G-CSF versus classic M-VAC in advanced urothelial tract tumours. Eur J Cancer 2006;42:50-54. [Crossref] [PubMed]
- von der Maase H, Sengelov L, Roberts JT, et al. Long-term survival results of a randomized trial comparing gemcitabine plus cisplatin, with methotrexate, vinblastine, doxorubicin, plus cisplatin in patients with bladder cancer. J Clin Oncol 2005;23:4602-8. [Crossref] [PubMed]
- Galsky MD, Hahn NM, Rosenberg J, et al. A consensus definition of patients with metastatic urothelial carcinoma who are unfit for cisplatin- based chemotherapy. Lancet Oncol 2011;12:211-4. [Crossref] [PubMed]
- Massari F, Di Nunno V, Cubelli M, et al. Immune checkpoint inhibitors for metastatic bladder cancer. Cancer Treat Rev 2018;64:11-20. [Crossref] [PubMed]
- Pignot G, Loriot Y, Kamat AM, et al. Effect of immunotherapy on local treatment of genitourinary malignancies. Eur Urol Oncol 2019;2:355-64. [Crossref] [PubMed]
- Powles T, Park SH, Voog E, et al. Maintenance avelumab + best supportive care (BSC) versus BSC alone after platinum-based first-line (1L) chemotherapy in advanced urothelial carcinoma (UC): JAVELIN Bladder 100 phase III interim analysis. J Clin Oncol 2020;38:18_suppl, LBA1-LBA1.
- Lattanzi M, Balar AV. Current status and future direction of immunotherapy in urothelial carcinoma. Curr Oncol Rep 2019;21:24. [Crossref] [PubMed]
- Kluth LA, Black PC, Bochner BH, et al. Prognostic and Prediction Tools in Bladder Cancer: A Comprehensive Review of the Literature. Eur Urol 2015;68:238-53. [Crossref] [PubMed]
- Bochner BH, Kattan MW, Vora KC, et al. Postoperative nomogram predicting risk of recurrence after radical cystectomy for bladder cancer. J Clin Oncol 2006;24:3967-72. [Crossref] [PubMed]
- Mollica V, Rizzo A, Montironi R, et al. Current Strategies and Novel Therapeutic Approaches for Metastatic Urothelial Carcinoma. Cancers 2020;12:E1449 [Crossref] [PubMed]
- Ma G, Yang X, Liang Y, et al. Precision medicine and bladder cancer heterogeneity. Bull Cancer 2018;105:925-31. [Crossref] [PubMed]
- Knowles MA, Hurst CD. Molecular biology of bladder cancer: new insights into pathogenesis and clinical diversity. Nat Rev Cancer 2015;15:25-41. [Crossref] [PubMed]
- Godwin JL, Hoffman-Censits J, Plimack E. Recent developments in the treatment of advanced bladder cancer. Urol Oncol 2018;36:109-14. [Crossref] [PubMed]
- Sjödahl G, Jackson CL, Bartlett JM, et al. Molecular profiling in muscle-invasive bladder cancer: more than the sum of its parts. J Pathol 2019;247:563-73. [Crossref] [PubMed]
- Vogelstein B, Papadopoulos N, Velculescu VE, et al. Cancer genome landscapes. Science 2013;339:1546-58. [Crossref] [PubMed]
- Morganti S, Tarantino P, Ferraro E, et al. Complexity of genome sequencing and reporting: Next generation sequencing (NGS) technologies and implementation of precision medicine in real life. Crit Rev Oncol Hematol 2019;133:171-82. [Crossref] [PubMed]
- Petersen BS, Fredrich B, Hoeppner MP, et al. Opportunities and challenges of whole-genome and -exome sequencing. BMC Genet 2017;18:14. [Crossref] [PubMed]
- Yohe S, Thyagarajan B. Review of Clinical Next-Generation Sequencing. Arch Pathol Lab Med 2017;141:1544-57. [Crossref] [PubMed]
- Nakagawa H, Fujita M. Whole genome sequencing analysis for cancer genomics and precision medicine. Cancer Sci 2018;109:513-22. [Crossref] [PubMed]
- Sabour L, Sabour M, Ghorbian S. Clinical Applications of Next-Generation Sequencing in Cancer Diagnosis. Pathol Oncol Res 2017;23:225-34. [Crossref] [PubMed]
- Nair M, Sandhu SS, Sharma AK. Cancer molecular markers: A guide to cancer detection and management. Semin Cancer Biol 2018;52:39-55. [Crossref] [PubMed]
- Fantini D, Meeks JJ. Genomic classification and risk stratification of bladder cancer. World J Urol 2019;37:1751-7. [Crossref] [PubMed]
- Agarwal N, Pal SK, Hahn AW, et al. Characterization of metastatic urothelial carcinoma via comprehensive genomic profiling of circulating tumor DNA. Cancer 2018;124:2115-24. [Crossref] [PubMed]
- Bellmunt J, von der Maase H, Mead GM, et al. Randomized phase III study comparing paclitaxel/cisplatin/gemcitabine and gemcitabine/cisplatin in patients with locally advanced or metastatic urothelial cancer without prior systemic therapy: EORTC Intergroup Study 30987. J Clin Oncol 2012;30:1107-13. [Crossref] [PubMed]
- Galsky MD, Hahn NM, Rosenberg J, et al. Treatment of patients with metastatic urothelial cancer “unfit” for Cisplatin-based chemotherapy. J Clin Oncol 2011;29:2432-8. [Crossref] [PubMed]
- Balar AV. Recent clinical trials explore immunotherapies for urothelial carcinoma. Oncology (Williston Park) 2019;33:132-6. [PubMed]
- Postow MA, Sidlow R, Hellman MD. Immune-related adverse events associated with immune checkpoint blockade. N Engl J Med 2018;378:158e68.
- Wang PF, Chen Y, Song SY, et al. Immune-related adverse events associated with anti-PD-1/PD-L1 treatment for malignancies: a meta-analysis. Front Pharmacol 2017;8:730. [Crossref] [PubMed]
- Wang DY, Salem JE, Cohen JV, et al. Fatal Toxic Effects Associated With Immune Checkpoint Inhibitors: A Systematic Review and Meta-analysis. JAMA Oncol 2018;4:1721-8. [Crossref] [PubMed]
- Belum VR, Benhuri B, Postow MA, et al. Characterisation and management of dermatologic adverse events to agents targeting the PD-1 receptor. Eur J Cancer 2016;60:12-25. [Crossref] [PubMed]
- Nadal R, Bellmunt J. Management of metastatic bladder cancer. Cancer Treat Rev 2019;76:10-21. [Crossref] [PubMed]
- Damrauer JS, Hoadley KA, Chism DD, et al. Intrinsic subtypes of high-grade bladder cancer reflect the hallmarks of breast cancer biology. Proc Natl Acad Sci U S A 2014;111:3110-5. [Crossref] [PubMed]
- Choi W, Porten S, Kim S, et al. Identification of distinct basal and luminal subtypes of muscle-invasive bladder cancer with different sensitivities to frontline chemotherapy. Cancer Cell 2014;25:152-65. [Crossref] [PubMed]
- Cancer Genome Atlas Research Network. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 2014;507:315-22. [Crossref] [PubMed]
- Robertson AG, Kim J, Al-Ahmadie H, et al. Comprehensive molecular characterization of muscle-invasive bladder cancer. Cell 2017;171:540-56.e25. [Crossref] [PubMed]
- Kamoun A, de Reynies A, Allory Y, et al. A Consensus Molecular Classification of Muscle-invasive Bladder Cancer. Eur Urol 2020;77:420-33. [Crossref] [PubMed]
- Alifrangis C, McGovern U, Freeman A, et al. Molecular and histopathology directed therapy for advanced bladder cancer. Nat Rev Urol 2019;16:465-83. [Crossref] [PubMed]
- Katoh M, Nakagama H. FGF receptors: cancer biology and therapeutics. Med Res Rev 2014;34:280-300. [Crossref] [PubMed]
- Billerey C, Chopin D, Aubriot-Lorton MH, et al. Frequent FGFR3 mutations in papillary non-invasive bladder (pTa) tumors. Am J Pathol 2001;158:1955-9. [Crossref] [PubMed]
- van Rhijn BW, Lurkin I, Radvanyi F, et al. The fibroblast growth factor receptor 3 (FGFR3) mutation is a strong indicator of superficial bladder cancer with low recurrence rate. Cancer Res 2001;61:1265-8. [PubMed]
- Cheng T, Roth B, Choi W, et al. Fibroblast growth factor receptors-1 and -3 play distinct roles in the regulation of bladder cancer growth and metastasis: implications for therapeutic targeting. PLoS One 2013;8:e57284 [Crossref] [PubMed]
- Milowsky MI, Dittrich C, Duran I, et al. Phase 2 trial of dovitinib in patients with progressive FGFR3-mutated or FGFR3 wild-type advanced urothelial carcinoma. Eur J Cancer 2014;50:3145-52. [Crossref] [PubMed]
- Rosenberg JE, Hoffman-Censits J, Powles T, et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet 2016;387:1909-20. [Crossref] [PubMed]
- Galsky MD, Saci A, Szabo PM, et al. Impact of tumor mutation burden on nivolumab efficacy in second-line urothelial carcinoma patients: exploratory analysis of the phase II checkmate 275 study. Ann Oncol 2017;28:v295-v329. [Crossref]
- Di Nunno V, De Luca E, Buttigliero C, et al. Immune-checkpoint inhibitors in previously treated patients with advanced or metastatic urothelial carcinoma: a systematic review and meta-analysis. Crit Rev Oncol Hematol 2018;129:124-32. [Crossref] [PubMed]
- Schwamborn K, Knüchel R, Hartmann A, et al. Comparability-of-programmed-death-ligand-1-PD-L1-expres- sion-on-tumor-infiltrating-immune-cells-IC-and-tumor-cells- TC-in-advanced-urothelial-bladder-cancer-UBC-using-clini- cally-relevant-immunohistochemistry-IHC-assays. Ann Oncol 2017;28:v403-27.
- Rizvi NA, Hellmann MD, Snyder A, et al. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 2015;348:124-8. [Crossref] [PubMed]
- Patel SP, Kurzrock R. PD-L1 Expression as a Predictive Biomarker in Cancer Immunotherapy. Mol Cancer Ther 2015;14:847-56. [Crossref] [PubMed]
- Gibney GT, Weiner LM, Atkins MB. Predictive biomarkers for checkpoint inhibitor- based immunotherapy. Lancet Oncol 2016;17:e542-51. [Crossref] [PubMed]
- Colli LM, Machiela MJ, Myers TA, et al. Burden of nonsynonymous mutations among TCGA cancers and candidate immune checkpoint inhibitor responses. Cancer Res 2016;76:3767-72. [Crossref] [PubMed]
- Lerner SP, McConkey DJ, Hoadley KA, et al. Bladder cancer molecular taxonomy: summary from a consensus meeting. Bladder Cancer 2016;2:37-47. [Crossref] [PubMed]
- Gajewski TF, Schreiber H, Fu YX. Innate and adaptive immune cells in the tumor microenvironment. Nat Immunol 2013;14:1014-22. [Crossref] [PubMed]
- Ikeda H, Old LJ, Schreiber RD. The roles of IFNγ in protection against tumor development and cancer immunoediting. Cytokine Growth Factor Rev 2002;13:95-109. [Crossref] [PubMed]
- Sharma P, Allison JP. The future of immune checkpoint therapy. Science 2015;348:56-61. [Crossref] [PubMed]
- Jamieson NB, Maker AV. Gene-expression profiling to predict responsiveness to immunotherapy. Cancer Gene Ther 2017;24:134-40. [Crossref] [PubMed]
- Wood RD, Mitchell M, Lindahl T. Human DNA repair genes, 2005. Mutat Res 2005;577:275-83. [Crossref] [PubMed]
- Turner N, Tutt A, Ashworth A. Hallmarks of 'BRCAness' in sporadic cancers. Nat Rev Cancer 2004;4:814-9. [Crossref] [PubMed]
- Urun Y, Leow JJ, Fay AP, et al. ERCC1 as a prog- nostic factor for survival in patients with advanced urothelial cancer treated with platinum based chemotherapy: a systematic review and meta-analysis. Crit Rev Oncol Hematol 2017;120:120-6. [Crossref] [PubMed]
- De Vos M, Schreiber V, Dantzer F. The diverse roles and clinical relevance of PARPs in DNA damage repair: current state of the art. Biochem Pharmacol 2012;84:137-46. [Crossref] [PubMed]
- Eustermann S, Wu WF, Langelier MF, et al. Structural Basis of Detection and Signaling of DNA Single-Strand Breaks by Human PARP-1. Mol Cell 2015;60:742-54. [Crossref] [PubMed]
- Teo MY, Bambury RM, Zabor EC, et al. DNA damage response and repair gene alterations are associated with improved survival in patients with platinum-treated advanced urothelial carcinoma. Clin Cancer Res 2017;23:3610-8. [Crossref] [PubMed]
- Sternberg CN, Yagoda A, Scher HI, et al. Cisplatin for advanced transitional cell carcinoma of the urothelium. Efficacy and patterns of response and relapse. Cancer 1989;64:2448-58. [Crossref] [PubMed]
- Mullane SA, Werner L, Guancial EA, et al. Expression Levels of DNA Damage Repair Proteins Are Associated With Overall Survival in Platinum-Treated Advanced Urothelial Carcinoma. Clin Genitourin Cancer 2016;14:352-9. [Crossref] [PubMed]
- Kondo T, Kanai M, Kou T, et al. Association between homologous recombination repair gene mutations and response to oxaliplatin in pancreatic cancer. Oncotarget 2018;9:19817-25. [Crossref] [PubMed]
- Ray Chaudhuri A, Callen E, Ding X, et al. Replication fork stability confers chemoresistance in BRCA-deficient cells. Nature 2016;535:382-7. [Crossref] [PubMed]
- Stühler V, Maas JM, Bochem J, et al. Molecular predictors of response to PD-1/PD-L1 inhibition in urothelial cancer. World J Urol 2019;37:1773-84. [Crossref] [PubMed]
- Teo MY, Seier K, Ostrovnaya I, et al. Alterations in DNA damage response and repair genes as potential marker of clinical benefit from PD-1/PD-L1 blockade in advanced urothelial cancers. J Clin Oncol 2018;36:1685-94. [Crossref] [PubMed]


