
Expression of G3BP1 in benign and malignant human prostate tissues
Introduction
Prostate cancer (PCa) is a leading type of cancer in men worldwide. According to the Global Burden of Disease Cancer, PCa is the highest incidence for men in 92 countries, and the leading cause of cancer deaths for men in 48 countries (1). In the United States, it is estimated that one in every nine men will be diagnosed with the disease during his lifetime (2).
Androgen receptor (AR) plays a central role in the initiation and progression of PCa. To date, androgen deprivation therapy (ADT) is the primary treatment for PCa to inhibits AR signaling. AR is the key driver of PCa progression and the main target for metastatic PCa treatment. Considering the reliance of PCa on AR, it is essential to identify AR-related genes involved in prostate tumor metastases.
RNA-binding proteins (RBPs) play an essential role in the regulation of gene expression (3). As a member of the RBP family, GTPase-activating protein SH3 domain-binding protein 1 (G3BP1) may be involved in tumorigenesis and development by interacting with its target mRNA (4). For example, some cancer related genes include c-MYC (5), CTNNB1 (6) and PMP22 (7) have been proved to be regulated by G3BP1. Early studies have shown that G3BP1 is overexpressed in many tumors (8-10). However, there are relatively few studies regarding G3BP1 in PCa.
In this study, we reported the first study of G3BP1 in PCa. G3BP1 was highly expressed in PCa, and its expression is associated with tumor malignancy. Furthermore, G3BP1 might be closely related to AR. We present the following article in accordance with the REMARK reporting checklist (available at http://dx.doi.org/10.21037/tau-20-1450).
Methods
UALCAN
UALCAN (http://ualcan.path.uab.edu/) is a comprehensive, user-friendly, and interactive web resource for analyzing cancer OMICS data (11). It built on PERL-CGI with high quality graphics using JavaScript and CSS. In this study, G3BP1 was submitted to UALCAN and its mRNA expression in PRAD was explored using TCGA PRAD samples (n=497). We further compared the expression across different tumor subgroups as defined by patient age, Gleason score and nodal metastasis status.
GEPIA2
GEPIA2 (http://gepia2.cancer-pku.cn/) is an updated version of GEPIA for analyzing the RNA sequencing expression data of 9,736 tumors and 8,587 normal samples from the TCGA and the GTEx projects, using a standard processing pipeline (12). We used GEPIA2 to verify the correlation between G3BP1 and AR mRNA expression. Three expression datasets (PRAD tumor, PRAD normal, prostate in GTEx) were used, and correlation coefficient (r) and P value were calculated by Pearson’s correlation analysis (R=0.64, P value =0)
Patients and tissue samples
Patients were identified from a population of men with PCa treated at the Second Hospital of Tianjin Medical University between 2016 and 2019. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by the Institutional Review Board of the Second Hospital of Tianjin Medical University (No. KY2019K036) and individual consent for this retrospective analysis was waived. These samples were used for immunohistochemistry (IHC) analysis. Clinical parameters including age, serum PSA, Gleason grade and metastasis states were collected. The clinical and pathological factors of all patients were shown in Table 1.
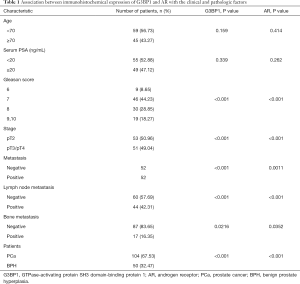
Full table
IHC techniques
Formalin-fixed, paraffin-embedded PCa and benign prostate hyperplasia (BPH) tissues were cut into 4 µm thickness sections. Tissue sections were deparaffinized with xylene, dehydrated in gradient ethanol, and then antigen retrieval was performed with microwave treatment in citrate buffer (pH =6.0) (Solarbio, China). Endogenous peroxidase activity was blocked in a 15 min treatment with 3% hydrogen peroxide. The sections were then incubated with anti-G3BP1 (Santa Cruz, USA, sc-98,561, 1:50) or anti-AR (Abcam, USA, ab108,341, 1:3,000) antibody diluted in PBS that contained 3% bovine serum albumin (Solarbio, China) overnight at 4 °C. After washing with PBS, the sections were incubated with secondary antibody (ZSGB-BIO, China) for 1 h at room temperature and visualized using the HRP DAB Detection Kit (ZSGB-BIO, China). Finally, sections were counterstained with hematoxylin.
For the scoring system, all slides immunostained with G3BP1 and AR antibody were scored by two independent researchers for the percentage and intensity of cells showing specific immunostaining signals. The percentage score grading criteria of 0–5 were as follows: 0, 0–1% of tumor cells positive; 1, 1–10% of tumor cells were positive; 2, 11–50% of cells were positive; 3, 50–75% of cells were positive; and 4, >75% of cells were positive. The intensity of immunostaining level was determined by the subjective visual scoring of the brown stain as follows: 0, no staining; 1, yellow-brown staining; 2, brown staining. The final score was calculated by combining the percentage and intensity scores and recorded as low [0–3], moderate [4–8] and high [9–12].
Cell culture and transfection
The human PCa cell lines LNCaP and the human embryonic kidney (HEK) 293T cells were acquired from American Type Culture Collection (ATCC, USA). The LNCaP cells were cultured in RPMI 1640 (Biological Industries, Israel) supplemented with 10% fetal bovine serum (Excell Bio, New Zealand) and 1% penicillin/streptomycin (Biological Industries, Israel) while HEK 293T cells were cultured in Dulbecco’s modified Eagle medium (DMEM) (Biological Industries, Israel), supplemented with 10% fetal bovine serum (Biological Industries, Israel). All cell lines were maintained at 37 °C and 5% CO2. To generate G3BP1 knockdown stable clones, HEK 293T cells were transfected with lentiviral vectors (Sigma, USA) at 70–80% cell density, including pLKO.1-Scr and pLKO.1-shG3BP1, together with lentivirus packaging plasmids (psPAX2 and pMD2.G) for 48 h using Lipofectamine 2000 (Invitrogen, USA). The lentivirus supernatant was collected and then added to the culture medium of prostate cancer cells for shRNA transduction. Two days after infection, stable clones were selected with 2 µg/mL puromycin (Sangon Biotech, China) for 10 days and puromycin-resistant cells were subsequently expanded with medium containing 0.5 µg/mL puromycin. The shG3BP1 sequence was: forward oligo, CCGGCGGGAATTTGTGAGACAGTATCGGGATCCAATACTGTCTCACAAATTCCCGTTTTTTG; reverse oligo, AATTCAAAAAACGGGAATTTGTGAGACAG-TAGGGTTATCCCGAGACTGTCTCACAAATTCCCG.
Western blotting
Cells were washed twice with ice-cold PBS, then lysed in SDS lysis buffer containing 1× protease inhibitor cocktail (Roche Applied Science, Germany). The total protein concentration in the cell lysate solution was then determined via the BCA protein assay (Thermo Fisher Scientific, USA). Protein samples (20 µg) were separated by electrophoresis on 10% SDS-PAGE and transferred to polyvinylidene difluoride membranes (Millipore, USA). After being blocked with 5% skim milk (BD Biosciences, USA) for 1 h, the membranes were incubated with specific primary antibodies; β-actin (Affinity Biosciences, USA, T0022; 1:4,000), G3BP1 (Santa Cruz, USA, sc-98,561; 1:1,000), AR (Abcam, USA, ab108,341, 1:3,000) overnight at 4 °C. After washing with TBST, the membranes were incubated with HRP-conjugated anti-mouse (Affinity Biosciences, USA, S0002, 1:3,000) or anti-rabbit (Affinity Biosciences, USA, S0001, 1:5,000) secondary antibodies for 1 h and visualized with an ECL system (Millipore, USA).
Statistical analysis
Because the data were not normally distributed, non-parametric statistical methods were used. For IHC, different groups were performed with Mann-Whitney test. Correlation between G3BP1 and AR was performed with Spearman’s rho correlation. For biochemical recurrence-free survival (BRFS), P values were calculated using the log-rank (Mantel-Cox) test. In all tests, P<0.05 was taken as the significance limit. All statistical tests were calculated using the SPSS version 21.0 (SPSS Inc.) and GraphPad Prism 8.0 (GraphPad).
Results
Database analysis of G3BP1 expression in PCa
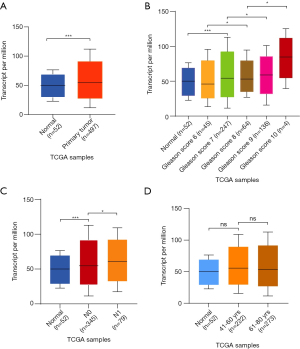
As a first step in analyzing the expression pattern of G3BP1 in PCa, we analyzed G3BP1 in PRAD patients by UALCAN. UALCAN analysis further showed that the RNA expression level of G3BP1 up-regulated significantly in primary tumor compared with the normal as shown in Figure 1A (P<0.001). Besides, it was found that G3BP1 expression was associated with Gleason score (Figure 1B) and nodal metastasis (Figure 1C), while there was no association with age (Figure 1D). A more detailed analysis between different Gleason score groups is presented in Table S1. Thus, the expression level of G3BP1 may be closely related to the occurrence and development of PCa.
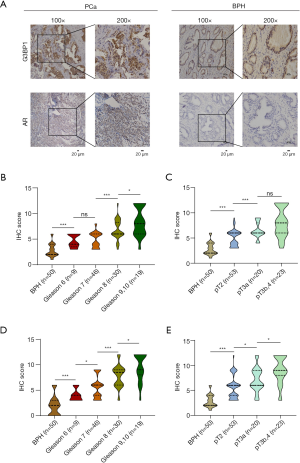
G3BP1 expression in clinical PCa cohorts
To further verify this result, we analyzed G3BP1 and AR expression by IHC staining in a panel of 104 PCa and 50 BPH samples. Similar to the results from RNA analyses, positive G3BP1 protein staining was strongly associated with PCa. Upregulation of G3BP1 expression was noticeable in PCa tissue compared with the BPH tissue (Figure 2A). Moreover, G3BP1 expression significantly correlated with both Gleason score (Figure 2B) and pT stage (Figure 2C), as well as AR (Figure 2D,E).
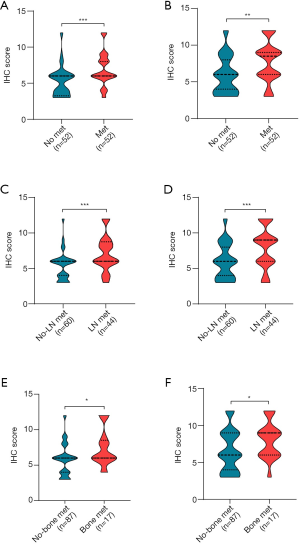
We also found that G3BP1 (Figure 3A) and AR (Figure 3B) were both associated with PCa metastasis, especially lymph node metastasis (Figure 3C,D). Bone is the most common site of metastasis in advanced PCa. Significance was also found in bone metastasis (Figure 3E,F). These results suggest that G3BP1 is closely related to metastasis.
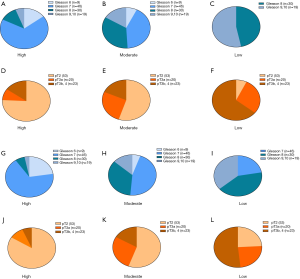
Next, we analyzed the expression of G3BP1 and AR in different subgroups. Patients were stratified according to the immunoreactive score as described above. Figure 4 showed that the expression of G3BP1 and AR was positively correlated with the malignancy of the tumor.
The relationship between G3BP1 and AR
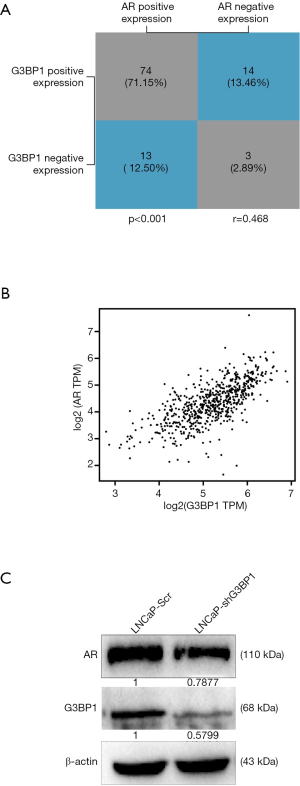
We inferred that there is some correlation between G3BP1 and AR. The results of IHC confirmed our conjecture (Spearman correlation r=0.468, P<0.001, Figure 5A). Further analyses of PCa patients from the GEPIA2 database revealed a positive correlation between G3BP1and AR mRNA (Person-correlation r=0.64) (Figure 5B). Next, we constructed the G3BP1 knockdown stable cell lines in LNCaP cells. We found that the downregulation of G3BP1 affected AR expression levels (Figure 5C). Although further work is required, these data indicate that G3BP1 is indeed closely related to AR.
Biochemical recurrence
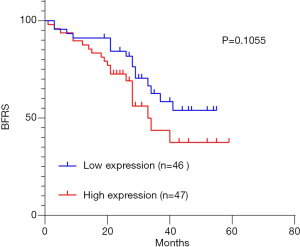
We next examined the significance of G3BP1 expression on biochemical recurrence by following up 93 patients (Figure 6). Recurrence-free survival was measured from the time of surgery to disease recurrence or death. We classified patients into high and low expression based on the median value. Although, the difference between low and high expression groups did not reach statistical significance, there was a strong trend towards a difference (P=0.1055). We suspected that this might be related to the small sample size and the short follow-up period.
Discussion
PCa has been the most common cancer among men in Western countries (13). Despite extensive studies, the fundamental mechanisms responsible for the development and progression of PCa have not yet been fully elucidated. Here we provide a new mechanistic link between G3BP1 and AR, which may be involved in the occurrence and development of PCa.
The primary objective of our study was to examine the expression of G3BP1 in PCa. In this work, we presented evidence that G3BP1 was frequently upregulated in PCa samples, in accordance with G3BP1 in other cancers. Statistical analysis revealed that high expression of G3BP1 was correlated with tumor malignancy. Indeed, our findings indicated that high G3BP1 expression might be predictive of biochemical recurrence. Meanwhile, metastatic PCa represents a small, but clinically significant, proportion of all PCa cases. The finding that G3BP1 was strongly associated with metastasis might provide a new target for metastatic PCa.
AR expression has been found in almost all primary and metastatic PCa, regardless of stage or grade (14). AR signaling remains active and supports the survival and growth of PCa cells. Understanding how AR-regulated genes promote tumor metastasis and targeting critical AR-regulated signaling pathways in lethal metastatic PCa may be a promising future for drug development. Given the change of AR expression following G3BP1 knockdown, we predict G3BP1 associates with AR and may serve as an upstream regulatory mechanism. Considering that G3BP1 belongs to RNA binding proteins and can influence mRNA stability (15,16), we suppose G3BP1 may regulate AR expression, but the concrete mechanism remains to be further studied.
One of the limitations of this study is that the median follow-up period of 27 months was relatively short in considering the long natural history of PCa. Moreover, as we have described, the sample size is relatively small. Future studies should increase the sample size and follow the patients during their entire lifespan to confirm our findings. On the other hand, the research evidence remains insufficient, and the proposed mechanisms are superficial, warranting deeper empirical studies.
Conclusions
In conclusion, we found that G3BP1 was frequently upregulated in PCa and closely related to AR expression and tumor metastasis. These results supply a potential target for the management of the PCa.
Acknowledgments
We would like to thank the researchers and study participants for their contributions. We are very thankful to Mr. Saravanan Subramanian for carefully reading and polishing the manuscript.
Funding: This work was supported by the National Natural Science Foundation of China (grant numbers 81772945, 81872078, and 21974094), the Natural Science Foundation of Tianjin (grant number 18JCYBJC26700 and 18JCYBJC25200), the Young Elite Scientists Sponsorship Program (grant number TJSQNTJ-2017-10), and National Training Program of innovation and Entrepreneurship for undergraduates (grant number 202010062044).
Footnote
Reporting Checklist: The authors have completed the REMARK reporting checklist. Available at http://dx.doi.org/10.21037/tau-20-1450
Data Sharing Statement: Available at http://dx.doi.org/10.21037/tau-20-1450
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tau-20-1450). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by the Institutional Review Board of the Second Hospital of Tianjin Medical University (No. KY2019K036) and individual consent for this retrospective analysis was waived.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Fitzmaurice C, Akinyemiju TF, Al Lami FH, et al. Global, regional, and national cancer incidence, mortality, years of life lost, years lived with disability, and disability-adjusted life-years for 29 cancer groups, 1990 to 2016: a systematic analysis for the Global Burden of Disease Study. JAMA Oncol 2018;4:1553-68. [Crossref] [PubMed]
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin 2019;69:7-34. [Crossref] [PubMed]
- Armakolas A, Dimakakos A, Loukogiannaki C, et al. IL-6 is associated to IGF-1Ec upregulation and Ec peptide secretion, from prostate tumors. Mol Med 2018;24:6. [Crossref] [PubMed]
- Alam U, Kennedy D. Rasputin a decade on and more promiscuous than ever? A review of G3BPs. Biochim Biophys Acta Mol Cell Res 2019;1866:360-70. [Crossref] [PubMed]
- Gallouzi IE, Parker F, Chebli K, et al. A novel phosphorylation-dependent RNase activity of GAP-SH3 binding protein: a potential link between signal transduction and RNA stability. Mol Cell Biol 1998;18:3956-65. [Crossref] [PubMed]
- Bikkavilli RK, Malbon CC. Arginine methylation of G3BP1 in response to Wnt3a regulates β-catenin mRNA. J Cell Sci 2011;124:2310-20. [Crossref] [PubMed]
- Winslow S, Leandersson K, Larsson C. Regulation of PMP22 mRNA by G3BP1 affects cell proliferation in breast cancer cells. Mol Cancer 2013;12:156. [Crossref] [PubMed]
- Xiong R, Gao JL, Yin T. G3BP1 activates the TGF-β/Smad signaling pathway to promote gastric cancer. Onco Targets Ther 2019;12:7149-56. [Crossref] [PubMed]
- Dou N, Chen J, Yu S, et al. G3BP1 contributes to tumor metastasis via upregulation of Slug expression in hepatocellular carcinoma. Am J Cancer Res 2016;6:2641-50. [PubMed]
- Zhang H, Ma Y, Zhang S, et al. Involvement of Ras GTPase-activating protein SH3 domain-binding protein 1 in the epithelial-to-mesenchymal transition-induced metastasis of breast cancer cells via the Smad signaling pathway. Oncotarget 2015;6:17039-53. [Crossref] [PubMed]
- Chandrashekar DS, Bashel B, Balasubramanya SAH, et al. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia 2017;19:649-58. [Crossref] [PubMed]
- Tang Z, Kang B, Li C, et al. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res 2019;47:W556-60 [Crossref] [PubMed]
- International Agency for Research on Cancer, World Health Organization. Cancer Today: Global Cancer Observatory. Accessed March 18, 2020. Available online: https://gco.iarc.fr/today/home
- Aurilio G, Cimadamore A, Mazzucchelli R, et al. Androgen receptor signaling pathway in prostate cancer: from genetics to clinical applications. Cells 2020;9:2653. [Crossref] [PubMed]
- Atlas R, Behar L, Elliott E, et al. The insulin-like growth factor mRNA binding-protein IMP-1 and the Ras-regulatory protein G3BP associate with tau mRNA and HuD protein in differentiated P19 neuronal cells. J Neurochem 2004;89:613-26. [Crossref] [PubMed]
- Lypowy J, Chen IY, Abdellatif M. An alliance between Ras GTPase-activating protein, filamin C, and Ras GTPase-activating protein SH3 domain-binding protein regulates myocyte growth. J Biol Chem 2005;280:25717-28. [Crossref] [PubMed]


