Long non-coding RNAs: new frontiers for advancing personalized cancer medicine in prostate cancer
The significance of long non-coding RNAs (lncRNAs) in prostate cancer (PCa) is rapidly gaining attention because of accumulating evidence that demonstrates their important biological roles in tumor development and progression, and their biomarker potential in the diagnosis and prognosis of this disease (1). The advances in next-generation sequencing and bioinformatics analysis have led to the discovery of numerous lncRNAs which have dysregulated expression in PCa (1,2). Some of these lncRNAs have been found to exhibit oncogenic function or act as tumor suppressors, while the functions of several others remain unknown.
We read with interest the recent study by White et al. (3). They performed an integrative analysis and found androgen-regulated prostate cancer associated transcript-14 (PCAT-14) as the most prevalent lncRNA that was overexpressed in prostate tumors relative to normal prostate. Lower PCAT-14 expression was associated with increasing Gleason score and poor outcome; i.e., higher probability of metastatic progression, PCa-specific mortality and lower overall survival in multiple independent cohorts and ethnicities. In contrast, in vitro experiments demonstrated that high PCAT-14 expression suppressed an aggressive phenotype via reduced cellular growth, migration, and invasion. Additionally, PCAT-14 expression was reduced in patients with metastatic PCa. PCAT-14 is a PCa- and lineage-specific lncRNA. It was previously identified as a marker of low grade and indolent disease by Shukla et al. (4), as their data suggested that PCAT-14 can be used as a diagnostic biomarker, and overexpression of PCAT-14 suppressed invasion in vitro. These findings were similar to the White et al. (3) study, which found that PCAT14 was highly upregulated in PCa and loss of PCAT-14 was predictive of aggressive disease and prognostic for poor outcome.
Taken together, and now validated in separate independent cohorts, the results of these two studies confirm that PCAT-14 represents a unique biomarker that can be used for PCa diagnosis (highly expressed in prostate tumors) and can act as both a prognostic (lower expression associated with poor outcome) and predictive biomarker (response to androgen deprivation therapy).
LncRNAs act broadly within gene networks to regulate the major pathways of cell growth, proliferation, migration, invasion, differentiation and survival (5). Recent studies have indicated that lncRNAs can mediate a “sponge” regulatory network (sequestering microRNAs) that can differentially affect the expression of many protein-coding PCa driver genes and key components of cancer-driving pathways during carcinogenesis (6). Moreover, some lncRNAs are linked to reactivation of the androgen receptor signaling axis and reprogramming of PCa cellular metabolism, and thus may be differentially expressed during various phases of tumor development and progression (2,7,8). This might explain why PCAT-14 expression initially increases during prostate tumorigenesis and then subsequently decreases in the metastatic setting.
Several PCa-specific or PCa-associated lncRNAs have been identified to date, but only a few have been validated in independent patient cohorts or approved for clinical practice. Prostate cancer antigen 3 (PCA3) is the most prominent and clinically-relevant PCa-associated lncRNA, which was initially described as a novel biomarker of PCa and subsequently developed as a promising urine test for this disease. The PCA3 lncRNA-based urine test is approved for the diagnosis of PCa and notably has shown better performance than prostate-specific antigen (PSA) in urinary detection of PCa (9). The major barriers to more widespread use of PCA3 are its inability to be used as a prognostic biomarker and contradictory studies on its value in the prediction of clinical-pathological features of PCa (10,11). The main clinically-relevant lncRNAs in PCa are summarized in Table 1.
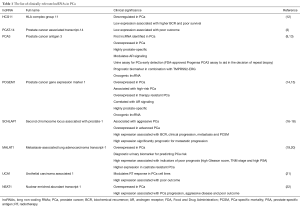
Full table
There is an urgent unmet need to develop sensitive and specific biomarkers for individualizing treatment recommendation in PCa. The promise of biomarkers to guide therapy is anticipated to extend to the radiotherapy (RT) setting. Recently, a systems biology-based radiosensitivity model [radiosensitivity index (RSI)] and a genomic-based clinical model [genomic-adjusted radiation dose (GARD)] have been developed which could help to predict intrinsic tumor radiosensitivity and personalize RT dose so that patients receive the optimum dose with an improved therapeutic ratio (23-25). Determining which patients harbor RT-resistant disease and are unlikely to derive a therapeutic benefit from RT will prevent overtreatment, thus removing the burden of unnecessary therapy and side-effects from patients and reduce costs to healthcare systems. Rather, these patients may benefit from radical prostatectomy or early integration of systemic therapies. To optimally integrate lncRNA-based biomarkers into the management of PCa patients being considered for RT, we need to identify the biomarkers that specifically predict RT response rather than those that are prognostic of outcome independent of treatment (26). For instance, we have recently discovered that the lncRNA urothelial carcinoma associated 1 (UCA1) mediates radiosensitivity in PCa cell lines and thus may be a promising biomarker to predict RT response in patients with PCa. UCA1 modulates radiosensitivity of PCa cells by impairing cell cycle progression, potentially through downregulation of the PI3K/Akt pathway (21).
Incorporating lncRNA biomarkers into standard risk stratification and as adjuncts to biomarkers that already exist could improve their specificity and sensitivity, help to precisely select aggressive from indolent cancer and optimize patient selection for definitive therapy. Combining these biomarkers with historical prognostic factors (PSA, Gleason score, clinical-pathological stage) would help to better predict treatment response and guide therapy decisions. The combination of different biomarkers together or with PSA (urinary TMPRSS2:ERG with urinary PCA3 and serum PSA) has been reported to provide high specificity and sensitivity compared to a single marker and increase the accuracy of prognostication (27). The detection of these lncRNAs is feasible in bodily fluids and may therefore be used as a liquid biopsy (9,19). Blood and urine-based biomarkers are ideal because they are minimally invasive and convenient for patients, can be readily monitored over the course of the disease and treatment, and are more representative of a patient’s entire PCa genome (compared to targeted biopsies). The identified biomarkers or combinations of markers require confirmation in large cohorts of patients to validate their specificity and sensitivity. These non-invasive lncRNAs could then be used to build biomarker signatures that serve to triage patients with aggressive disease for alternate or more intensive therapies and to identify a subset of patients for future biomarker-driven clinical trials. These strategies could potentially increase response rates as patients are triaged to the most appropriate treatment regimen and those patients who are unlikely to benefit are spared unnecessary side-effects of therapy.
LncRNAs also offer the potential to be a novel class of cancer therapeutic targets in the future. The strategies could be either to suppress oncogenic function or to activate tumor-suppressive activity of prostate-specific lncRNAs. To achieve this, further investigations are required to understand the functional role and molecular mechanisms of lncRNAs involved in PCa oncogenesis or tumor suppression, and characterize the critical mediators for selective cell killing.
Acknowledgements
Funding: AF Ghiam is supported by a Canadian Association of Radiation Oncology (CARO) Fellowship Award and University of Toronto Postgraduate Medical Education (PGME) Research Awards. SK Liu is a Movember Rising Star award recipient and is proudly funded by the Movember Foundation—Grants #RS2014-03 and #D2015-12, and through a Ministry of Research and Innovation Early Researcher Award (SK Liu). SK Liu and D Vesprini are also funded through the Prostate Cancer Fight Foundation—Telus Motorcycle Ride for Dad.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
References
- Ramalho-Carvalho J, Fromm B, Henrique R, et al. Deciphering the function of non-coding RNAs in prostate cancer. Cancer Metastasis Rev 2016;35:235-62. [Crossref] [PubMed]
- Guo H, Ahmed M, Zhang F, et al. Modulation of long noncoding RNAs by risk SNPs underlying genetic predispositions to prostate cancer. Nat Genet 2016;48:1142-50. [Crossref] [PubMed]
- White NM, Zhao SG, Zhang J, et al. Multi-institutional Analysis Shows that Low PCAT-14 Expression Associates with Poor Outcomes in Prostate Cancer. Eur Urol 2017;71:257-66. [Crossref] [PubMed]
- Shukla S, Zhang X, Niknafs YS, et al. Identification and Validation of PCAT14 as Prognostic Biomarker in Prostate Cancer. Neoplasia 2016;18:489-99. [Crossref] [PubMed]
- Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet 2009;10:155-9. [Crossref] [PubMed]
- Du Z, Sun T, Hacisuleyman E, et al. Integrative analyses reveal a long noncoding RNA-mediated sponge regulatory network in prostate cancer. Nat Commun 2016;7:10982. [Crossref] [PubMed]
- Weiss M, Plass C, Gerhauser C. Role of lncRNAs in prostate cancer development and progression. Biol Chem 2014;395:1275-90. [Crossref] [PubMed]
- Ma G, Tang M, Wu Y, et al. LncRNAs and miRNAs: potential biomarkers and therapeutic targets for prostate cancer. Am J Transl Res 2016;8:5141-50. [PubMed]
- de Kok JB, Verhaegh GW, Roelofs RW, et al. DD3(PCA3), a very sensitive and specific marker to detect prostate tumors. Cancer Res 2002;62:2695-8. [PubMed]
- Hendriks RJ, van Oort IM, Schalken JA. Blood-based and urinary prostate cancer biomarkers: a review and comparison of novel biomarkers for detection and treatment decisions. Prostate Cancer Prostatic Dis 2017;20:12-19. [Crossref] [PubMed]
- Hessels D, Schalken JA. The use of PCA3 in the diagnosis of prostate cancer. Nat Rev Urol 2009;6:255-61. [Crossref] [PubMed]
- Zhang Y, Zhang P, Wan X, et al. Downregulation of long non-coding RNA HCG11 predicts a poor prognosis in prostate cancer. Biomed Pharmacother 2016;83:936-941. [Crossref] [PubMed]
- Leyten GH, Hessels D, Jannink SA, et al. Prospective multicentre evaluation of PCA3 and TMPRSS2-ERG gene fusions as diagnostic and prognostic urinary biomarkers for prostate cancer. Eur Urol 2014;65:534-42. [Crossref] [PubMed]
- Srikantan V, Zou Z, Petrovics G, et al. PCGEM1, a prostate-specific gene, is overexpressed in prostate cancer. Proc Natl Acad Sci U S A 2000;97:12216-21. [Crossref] [PubMed]
- Petrovics G, Zhang W, Makarem M, et al. Elevated expression of PCGEM1, a prostate-specific gene with cell growth-promoting function, is associated with high-risk prostate cancer patients. Oncogene 2004;23:605-11. [Crossref] [PubMed]
- Prensner JR, Zhao S, Erho N, et al. RNA biomarkers associated with metastatic progression in prostate cancer: a multi-institutional high-throughput analysis of SChLAP1. Lancet Oncol 2014;15:1469-80. [Crossref] [PubMed]
- Prensner JR, Iyer MK, Sahu A, et al. The long noncoding RNA SChLAP1 promotes aggressive prostate cancer and antagonizes the SWI/SNF complex. Nat Genet 2013;45:1392-8. [Crossref] [PubMed]
- Mehra R, Udager AM, Ahearn TU, et al. Overexpression of the Long Non-coding RNA SChLAP1 Independently Predicts Lethal Prostate Cancer. Eur Urol 2016;70:549-52. [Crossref] [PubMed]
- Wang F, Ren S, Chen R, et al. Development and prospective multicenter evaluation of the long noncoding RNA MALAT-1 as a diagnostic urinary biomarker for prostate cancer. Oncotarget 2014;5:11091-102. [Crossref] [PubMed]
- Ren S, Liu Y, Xu W, et al. Long noncoding RNA MALAT-1 is a new potential therapeutic target for castration resistant prostate cancer. J Urol 2013;190:2278-87. [Crossref] [PubMed]
- Fotouhi Ghiam A, Taeb S, Huang X, et al. Long non-coding RNA urothelial carcinoma associated 1 (UCA1) mediates radiation response in prostate cancer. Oncotarget 2017;8:4668-89. [PubMed]
- Chakravarty D, Sboner A, Nair SS, et al. The oestrogen receptor alpha-regulated lncRNA NEAT1 is a critical modulator of prostate cancer. Nat Commun 2014;5:5383. [Crossref] [PubMed]
- Eschrich SA, Pramana J, Zhang H, et al. A gene expression model of intrinsic tumor radiosensitivity: prediction of response and prognosis after chemoradiation. Int J Radiat Oncol Biol Phys 2009;75:489-96. [Crossref] [PubMed]
- Eschrich SA, Fulp WJ, Pawitan Y, et al. Validation of a radiosensitivity molecular signature in breast cancer. Clin Cancer Res 2012;18:5134-43. [Crossref] [PubMed]
- Scott JG, Berglund A, Schell MJ, et al. A genome-based model for adjusting radiotherapy dose (GARD): a retrospective, cohort-based study. Lancet Oncol 2017;18:202-11. [Crossref] [PubMed]
- Korpela E, Vesprini D, Liu SK. MicroRNA in radiotherapy: miRage or miRador? Br J Cancer 2015;112:777-82. [Crossref] [PubMed]
- Sanguedolce F, Cormio A, Brunelli M, et al. Urine TMPRSS2: ERG Fusion Transcript as a Biomarker for Prostate Cancer: Literature Review. Clin Genitourin Cancer 2016;14:117-21. [Crossref] [PubMed]


